New & Noteworthy
Yeast Show No Fear of Commitment
March 11, 2022
Meiosis in budding yeast is typically induced by starvation, with a signal roughly translated as “make spores quick because things are terrible right now.” When the outside food supply dries up, meiosis gets initiated in the nucleus of the yeast cell. If food is resupplied early enough, meiosis can pause and mitosis—a normal cell division—can be undergone instead, a process called “return to growth” (RTG). There’s a sharp point-of-no-return in mid-prometaphase, though, after which no amount of food will make cells go back to mitosis—they’ve committed to meiosis.

The components involved in this commitment to meiosis have been poorly understood. It was shown by several studies that the transcription factor Ndt80p is required at high levels to establish meiotic commitment by inducing middle meiosis genes. When levels of Ndt80p are low, in contrast, cells show a commitment defect in which they go back to mitosis even after initiating meiosis, leading to defective polyploid germ cells with multiple nuclei.

Gavade et al., in a study that just came out in Current Biology, discover six novel regulators of meiotic commitment. The authors first identified proteins whose overexpression reversed the commitment defects of a low-Ndt80 strain. The pool of potential regulators was narrowed by looking at mutants of the genes identified in the overexpression screen to see if decreased levels would cause commitment defects. They found BCY1 (involved in nutrient sensing), IME1 (a meiosis-specific kinase), CDC5 (Polo kinase, with related roles in mitosis), BMH1 and BMH2 (14-3-3- proteins involved in numerous types of signaling), and PES4 (an RNA-binding protein with an uncharacterized role in cell cycle control) to all have roles in meiotic commitment. The authors further showed that Bmh1p and Bmh2p are direct regulators of both Ndt80p and Cdc5p, the former by protein stability and the latter by protein activation.

This intriguing study is an excellent example of how complex processes—such as the effect of food on sexual reproduction—can be teased apart in yeast. A relatively simple screen was able to identify proteins not formerly known to have a role in meiotic commitment, and it will be exciting to see if these proteins have orthologs in mammals that play related roles.
Categories: Research Spotlight
Tags: meiosis, meiotic commitment, response to starvation, response to stress, Saccharomyces cerevisiae
Phosphosite Library Pinpoints Link from Stress to Chromatin
March 05, 2022
The “language” of histone methylation has been a subject of intensive study due to numerous diseases and disorders linked to faulty methylation patterns. Methylation patterns are “written” by enzymes in response to signals and then “read” by effector proteins recognizing methyl residues on highly specific lysine residues, leading to either large- or small-scale alterations in the transcriptional state of chromatin. In response to the cellular environment, signals are sent for the opposing processes of “writing” and “erasing” methylation. Conserved methyltransferase enzymes are the “writers” and demethylase enzymes are the “erasers,” with the activity of each regulated by cellular signals in ways that are poorly understood.
Whereas humans have 35 writers and 23 erasers, yeast has only four of each. Given orthology within each class of writers and erasers (as defined by the particular lysines methylated or demethylated), this makes yeast a perfect model system for digging into the links that connect cellular signals to specific methylation patterns on chromatin.

In a recent study in the Journal of Molecular Biology, Separovich et al. describe a systematic phosphosite mutant library that allowed the identification of key phosphorylated residues transducing cellular signals onto a writer/eraser pair. In response to environmental stress, Set2p methylates lysine 36 on histone H3 while Jhd1p opposes this action by demethylation. Using AlphaFold, they modeled the relationship between the specific phosphorylated residues and showed the key regulator of methylation activity (T127 on Set2p) is spatially proximal to the target lysine residue in the histone.
Upon analysis of differential expression for the sets of phosphonull and phosphomimetic mutants, they showed the proteins most affected by histone methylation clustered into GO categories consistent with cellular response to stress, e.g. ion membrane transport, lipid biosynthesis, ergosterol biosynthesis, and protein mannosylation.
While the kinase(s) responsible for phosphorylating the writer/eraser pair have yet to be identified, there are good candidates to test in yeast. The identification of the yeast players in signal transduction from environment to chromatin will undoubtedly be of use to those studying the much more complex system in humans.
Categories: Research Spotlight
Tags: chromatin methylation, chromatin remodeling, histone methylation, Saccharomyces cerevisiae, yeast model for human disease
Green Monster Evolution to Hulk
February 25, 2022
A clever new study has used a modified yeast strain “ABC16-Green Monster” with fewer export pumps (i.e. more susceptible to drugs) to exert powerful selection pressure on yeast cells. By exposing this strain to multiple libraries of chemical compounds with potential use as antifungal agents, Ottilie et al. in a recent issue of Communications Biology identified an evolved set of 25 genes with frequent mutations.

In comparing the set of mutations to those frequently found upon extended growth without selection, they noted that intergenic mutations were comparatively rare, presumably due to the heavy selection pressure for functional resistance. For another confirmation of the screen’s effectiveness in isolating useful variants, i.e. not passenger mutations, they introduced 61 of the altered alleles back into the unevolved strain by CRISPR/Cas9 integration and verified that 45 variants across 37 genes restored resistance.
Looking more closely at the mechanisms of action for the observed resistances, they noted mutations clustered in the active sites of target molecules for which the target was previously known. For example, the antifungal drug tavaborole (a type of benzoxaborole) led to identification of four active-site mutations in Cdc60p (the yeast leucyl-tRNA synthetase) that are predicted to interfere with binding to the drug. In a related example, the chemotherapy drug camptothecin inhibits topoisomerase and they isolated two mutations in TOP1 that would be expected to fall in the binding pocket.
Their isolation of mutations in TOR2 and FPR1 (both associated with TOR signaling) in strains with evolved resistance to rapamycin reproduce the findings of a related study looking at rapamycin resistance in yeast. In fact, both studies identified mutations in the same S1975 residue of TOR2. In other interesting findings, they isolated mutations leading to yeast resistance to drugs used typically against soil-transmitted helminths (worms), trypanosomatid parasites, and malarial Plasmodium. Each mutant variant in yeast affords insight into how these pathogens might likewise evolve resistance or could be alternately targeted.

The set of identified mutant genes was highly enriched for transcription factors and among these were two—YRR1 and its paralog YRM1—that together mediated resistance for nearly 25% of the compounds tested. These two zinc transcription factors had previously been shown to activate genes involved in multidrug resistance. In this study, they were mutated 100 times in screens against 19 diverse chemical compounds. Interestingly, deletion of these genes does not confer resistance and the sum of the data suggests the screen identified gain-of-function alleles. In support of this idea, integration of the L611F allele of YRR1 into a susceptible strain by CRISPR/Cas9 reconstructed resistance to a suite of compounds. The researchers hypothesize that modified proteins lead to constitutive activation of genes aiding resistance.
In sum, this study demonstrates the awesome power of yeast genetics (#APOYG) for revealing insight into the molecular underpinnings of medical chemistry. Studies like this one provide not only data but intriguing clues about where to look next—most of which will be easy to test in yeast.
Categories: Research Spotlight
Tags: Drug resistance, Multidrug resistance, Saccharomyces cerevisiae, yeast model for human disease
C-Circles, an ALT Plan for Telomere Restoration
February 18, 2022
The telomerase ribonucleoprotein complex is the primary means by which yeast cells maintain telomeres. However, it turns out that cells lacking functional telomerase have a backup plan to restore telomere length by “alternative lengthening of telomeres” (ALT). ALT employs recombination via extrachromosomal telomere elements called C-circles. In a process for which the reasons remain unclear, C-circles get paired with eroded telomeres at the nuclear pore complex on the nuclear membrane. This pairing requires the SAGA/TREX2 complex and, once paired, the recombination between C-circles and telomeres appears to be effected by Rad59p, the paralog of Rad52p.

This interesting model is described in a recent paper in The EMBO Journal, in which Aguilera et al. adapt a method developed in human cancer studies to detect ALT and C-circles in yeast. In humans, ~10% of cancers depend on ALT for unchecked growth. In yeast, cells with ALT were able to be detected as survivors among telomerase mutant (est2∆) cells.
As other types of extrachromosomal DNA circles were previously reported to associate with the nuclear pore complex, the authors addressed the possibility that C-circles bind the NPC and demonstrated it clearly. They also showed the circles interact with the SAGA/TREX2 complex, which favors telomere recombination.
The novel finding that ALT in yeast so closely mirrors that of some human cancer cells is a boon to study of these cancers. The ability to develop ALT inhibitors in yeast would provide a new set of potential anticancer therapies, making this an ideal model system.
Categories: Research Spotlight
Tags: cancer, cell aging, Saccharomyces cerevisiae, senescence, telomeres, yeast model for human disease
Memories of Deceptive Courtship
February 10, 2022
Yeast courtship provides an excellent model for how a simple organism manages to remember past events. Recent yeast studies reveal how this memory can involve prion-like proteins that build up in a cell and persist as a form of memory. In a recent online issue of Current Biology, Lau et al. show how Whi3p self-templates into large assemblies upon perception of “deceptive courtship,” i.e. when pheromone is perceived but no mating partner appears. The super-assembly state allows escape from the G1 arrest triggered by potential courtship, but also prevents any further response to pheromone. While this pheromone refractory state is stable for the remainder of this mother cell’s life, daughter cells do not inherit this state and are fully pheromone responsive.

The organization of domains in Whi3p and the ability to self-template into large assemblies are features shared with prions. The lack of infectious heritability, though, distinguishes Whi3p as a mnemon rather than a prion. Fascinatingly, Lau et al. show that assembled Whi3p becomes heritable in cells with defective diffusion barriers, where the physical barriers at the bud neck normally restrict the super-assembled protein to the mother cell.

In studies of diffusion barrier disruption, the authors observed that the main super-assembly of Whi3p remains in the mother cell, while the daughter cells get “seeds” of Whi3mnem upon which to assemble further. They note that transmission of seeds is most prevalent in the first few divisions following escape from pheromone arrest, which suggests there is a limiting factor to this diffusion. They propose a model in which the assembled Whi3mnem “matures” to a form that no longer propagates seeds to daughter cells, even in the absence of a barrier.
Thus, remarkably, prion-like behavior appears to be one of the most rudimentary forms of memory, and may have implication for understanding cell memory in higher organisms.
Categories: Research Spotlight
Tags: mnemons, prion-like proteins, Saccharomyces cerevisiae, yeast cell memory, yeast mating
New ncRNA Represses Zinc Regulator
February 04, 2022
In a twist to an established story, the termination of noncoding RNAs by the NNS complex (NRD1 snoRNA termination complex) appears to be dependent on the phosphorylation of a regulatory component. In the latest issue of Nucleic Acids Research, Haidara et al. show how the NNS-complex component Sen1p acts to repress transcription of the zinc master regular ZAP1 when Sen1p is phosphorylated, which appears to happen in response to excess zinc.
The NNS complex had previously been shown to terminate transcription of PHO84 via antisense RNA. In this current paper, the authors identify the previously unannotated noncoding RNA ZRN1 as lying directly upstream of ZAP1 and, when transcribed without termination, repressing the downstream gene. Termination of ZRN1 transcription by dephosphophorylated Sen1p derepresses ZAP1 mRNA levels via removal of the interfering RNA. As evidence of this relationship, a Sen1p phospho-mimetic mutation (T1623E) results in stable repression of ZAP1 transcription.

The model proposed is that zinc excess leads to phosphorylation of Sen1p by an unidentified kinase, which then causes the level of ZRN1 transcript to increase because termination is impaired. This in turn represses ZAP1 mRNA levels by interference, thereby repressing genes responsible for increased zinc uptake and storage.
Interestingly, the same system (i.e. Sen1p as a component of the NNS complex) represses PHO84 expression via interference, and PHO84 encodes a low-affinity Zn transporter that also contributes to zinc homeostasis. Might RNA interference play an expanded regulatory role over what is currently known?
Categories: Research Spotlight
Tags: noncoding RNA, Saccharomyces cerevisiae, transcription, zinc homeostasis
New Understanding of ER Membrane Expansion
January 28, 2022
How lipid metabolism gets coordinated with membrane growth in response to environment has remained an unexplained phenomenon. A recent paper explains key components in how cells manage this response.

Papagiannidis et al. demonstrate a key regulatory step via the transmembrane protein Ice2p. Ice2p affects negative regulation of Pah1p (called lipin in metazoa) by inhibiting the Nem1-Spo7 phosphatase complex, which would normally activate Pah1p by dephosphorylation. The study shows that Ice2p also plays a role in transcriptional regulation of lipid synthesis genes, thereby providing a key pivot in the cellular need to make more membrane in response to ER stress.
Categories: Paper of the Week
Tags: endoplasmic reticulum, lipid regulation, Membrane biogenesis, Saccharomyces cerevisiae
Predicted 3D Structures of Yeast Complexes
January 20, 2022
In an exciting new paper, Humphreys et al. describe the use of deep-learning-based algorithms to predict structures of not only single proteins, but assemblies of proteins. The team used rapid RoseTTAFold combined with the more accurate AlphaFold to build structural models for 106 previously unidentified protein assemblies and 806 complexes that had not been structurally characterized. The complexes have up to five subunits and are involved in numerous critical roles in cell biology.

Go look for your own proteins of interest at the ModelArchive and search in the Home page. Also find the link on the resources section of the SGD Interaction and Protein pages.
Categories: Announcements, Data updates, Paper of the Week
Tags: protein complex, Saccharomyces cerevisiae, yeast protein assembly
SGD Newsletter, Fall 2021
December 14, 2021
About this newsletter:
This is the Fall 2021 issue of the SGD newsletter. The goal of this newsletter is to inform our users about new features in SGD and to foster communication within the yeast community. You can view this newsletter as well as previous newsletters on our Community Wiki.
Contents
- 1Protein Complex Page Updates
- 2Nomenclature Updates
- 3New links to AlphaFold 3D Predicted Protein Structure Database
- 4DIOPT Orthologs and new queries in YeastMine
- 5Alliance of Genome Resources – latest release
- 6Upcoming Conferences and Courses
- 7Gene Ontology Consortium Fall 2021 Meeting
- 8Happy Holidays from SGD!
Protein Complex Page Updates
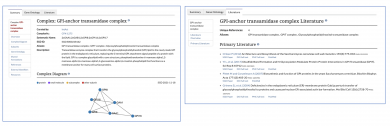
SGD has made recent updates to our protein complex pages to improve clarity and ease of use. The new pages for each complex will have the same format as gene pages, with tabs across the top for each category of information, including a Summary page, a Gene Ontology page, and a Literature page. Just as we do for all of your favorite genes, Gene Ontology and Literature curation for complexes will be ongoing.
If you have any questions or feedback about the updates to our complex pages, please do not hesitate to contact us at any time.
Nomenclature Updates
SGD has long been the keeper of the official Saccharomyces cerevisiae gene nomenclature. Robert Mortimer handed over this responsibility to SGD in 1993 after maintaining the yeast genetic map and gene nomenclature for 30 years.
The accepted format for gene names in S. cerevisiae comprises three uppercase letters followed by a number. The letters typically signify a phrase (referred to as the “Name Description” in SGD) that provides information about a function, mutant phenotype, or process related to that gene, for example “ADE” for “ADEnine biosynthesis” or “CDC” for “Cell Division Cycle”. Gene names for many types of chromosomal features follow this basic format regardless of the type of feature named, whether an ORF, a tRNA, another type of non-coding RNA, an ARS, or a genetic locus. Some S. cerevisiae gene names that pre-date the current nomenclature standards do not conform to this format, such as MRLP38, RPL1A, and OM45.
A few historical gene names predate both the nomenclature standards and the database, and were less computer-friendly than more recent gene names, due to the presence of punctuation. SGD recently updated these gene names to be consistent with current standards and to be more software-friendly by removing punctuation. The old names for these four genes have been retained as aliases.
Legacy gene names
| ORF | Old gene name | New gene name |
|---|---|---|
| YGL234W | ADE5,7 | ADE57 |
| YER069W | ARG5,6 | ARG56 |
| YBR208C | DUR1,2 | DUR12 |
| YIL154C | IMP2′ | IMP21 |
New systematic nomenclature for yeast genes not in the reference genome
For many years, a widely adopted systematic nomenclature has existed for yeast protein-coding genes, or ORFs, as many yeast researchers call them. Readers of the last SGD newsletter will recall that, earlier this year, SGD adopted a new systematic nomenclature for the entire annotated complement of ncRNAs.
We have just put into place a new systematic nomenclature for S. cerevisiae genes that are not found in the reference genome of strain S288C (“non-reference” genes). This new systematic nomenclature is similar to, but distinct from, that used for ORFs and that used for ncRNAs. Non-reference genes are designated by a symbol consisting of three uppercase letters and a four-digit number, as follows: Y for “Yeast”, SC for “Saccharomyces cerevisiae”, and a four-digit number corresponding to the sequential order in which the gene was added to SGD. We currently have 55 of these genes in SGD, some of which are old favorites like MAL21/YSC0004 and MATA/YSC0046, while others are more recent additions like XDH1/YSC0051. Going forward, as evidence is published pointing to other S. cerevisiae genes not present in the S288C reference genome, they will be added to the annotation using the next sequential number available. We already have 15 more of these YSC0000 names reserved by researchers and awaiting publication.
If you have some non-reference genes for which these names would be appropriate, please let us know!
New links to AlphaFold 3D Predicted Protein Structure Database
Would you like to see the shape of your protein?
SGD now contains links to AlphaFold in the Resources sections of the Summary, Protein, and Homology pages for every gene.
- The links through SGD give quick access to EMBL’s European Bioinformatics Institute (EMBL-EBI), which offers a new, highly accurate tool for predicting protein structure with speed and clarity.
- Given a peptide sequence for an uncharacterized protein, AlphaFold will model predicted domains and provide relative confidence levels for each portion of the prediction.
- The predicted domains can then be compared to known protein structures (using a tool such as PDBeFold) to seek matches to characterized protein families.
- Whether or not a family is identified, the comparison will yield clues to protein function to help design the next experiments.
DIOPT Orthologs and New Queries in YeastMine
We recently replaced HomoloGene, Ensembl, TreeFam and PANTHER homology datasets in YeastMine with homology data from DIOPT (DRSC integrative ortholog prediction tool). DIOPT integrates orthology predictions from multiple sources, including HomoloGene, Ensembl, TreeFam, and PANTHER. Using the Gene->Non-fungal and S. cerevisiae Homologs pre-generated query, you can look for DIOPT homologs for a single or multiple yeast genes. The results table provides identifiers and standard names for the yeast and homologous genes, as well as organism and predictive score information. As with other YeastMine templates, results can be saved as lists and analyzed further.
Pre-generated queries for human homolog(s) of your favorite yeast gene and their corresponding disease associations remain largely unchanged. You can begin with your favorite human gene or disease keyword and retrieve the yeast counterparts of the relevant gene(s). As an example, you can search for the S. cerevisiae homologs of all human genes associated with disorders that contain the keyword “diabetes” (view search). The results table provides identifiers and standard names for the yeast and human genes, OMIM gene and disease identifiers and name, as well as predictive algorithm sources and scores.
Alliance of Genome Resources – Recent Release
The Alliance of Genome Resources, a collaborative effort from SGD and other model organism databases (MOD), released version 4.1 this past August. Notable improvements and new features include:
- Human and model organism high throughput (HTP) variant data
- Human variants are imported from Ensembl
- Model organism HTP variants are submitted by Alliance members (FlyBase, RGD, SGD, Wormbase) or imported from EVA (MGI and ZFIN).
- Added HTP variants to the Alleles and Variants table on gene pages (e.g. rat Lepr Gene page) and to the table on the Alleles and Variants Details page (e.g. rat Lepr Alleles and Variants Details.
- Created a report page for Human and model organism HTP variants (e.g. human variant rs1041354454).
- Expanded Allele Category in search to “Allele/Variant” and added a search for HTP variants.
- On Gene Pages, a new Pathways widget displays via tabs:
- Reactome models of pathways for human gene products as well as inferred pathways for model organism genes based on orthology to human genes.
- Reactome reactions for gene products (e.g. human TP53 Gene page)
- Gene Ontology Causal Activity Models (GO-CAMs). These provide a framework to represent a biological system by linking together multiple GO annotations. PMID:31548717 (e.g. worm nsy-1 Gene page).
- Experimental conditions are include for Disease and Phenotype data in tables on Gene, Allele, and Disease pages (e.g. zebrafish scn1lab Gene page).
- AllianceMine added Orthologs, and Allele and Variants (low throughput) data types to this release. You can now query for these data types via pre-made template queries.
- The Alliance Community Forum is released. The Forum permits discussions across six model organism communities—flies, mice, yeast, rats, worms, and zebrafish. More details will follow.
Upcoming Conferences and Courses
- Fungal Genetics – the premier meeting for the international community of fungal geneticists
- Asilomar Conference Grounds, Pacific Grove, California (and Online)
- March 15 – 20, 2022
- 36th International Specialised Symposium on Yeasts (ISSY36) – Yeast Sea to Sky – Yeast in the Genomics Era
- University of British Columbia, Vancouver
- July 12 – 16, 2022
- CSHL Yeast Genetics & Genomics – modern, intensive laboratory course that teaches students full repertoire of genetic and genomic approaches
- Cold Spring Harbor Laboratory, NY
- July 26 – August 15, 2022
- Yeast Genetics Meeting – the premier meeting for students, postdoctoral scholars, research staff, and principal investigators studying various aspects of eukaryotic biology in yeast
- University of California, Los Angeles
- August 17 – 21, 2022
Gene Ontology Consortium Fall 2021 Meeting
From October 12-14, SGD biocurators attended the Gene Ontology Consortium’s Fall Meeting with participants from around the world. The goal of these meetings is to bring together data scientists with diverse backgrounds (curators, programmers, etc.) for lively discussions regarding how to better capture, curate, analyze, and serve data to researchers, educators, students, and other life science professionals. Our goal in participating in these meetings each year is to find ways to make SGD even better for you!
Discussion topics included, but were not limited to:
- LitSuggest – web-based system for biomedical literature recommendation and curation
- ECO, Evidence and Conclusions Ontology – terms used to describe types of evidence and assertion methods
- PAINT, Phylogenetic Annotation and INference Tool from PANTHER – orthology between reference genome genes and human disease genes
Happy Holidays from SGD!
We know that 2021 has been another challenging year for everyone. Our thoughts go out to all those who have been impacted by recent events. We wish you and your family, friends, and lab mates the best during the upcoming holidays.
Stanford University will be closed for two weeks starting December 20, and will reopen on January 3rd, 2022. Although SGD staff members will be taking time off, the website will be up and running throughout the winter break, and we will resume responding to user requests and questions in the new year.
Categories: Newsletter
Tags: Newsletter, Saccharomyces cerevisiae
Protein Complex Page Updates
December 01, 2021
SGD has updated our protein complex pages to have the same format as gene pages, with tabs across the top for each category of information, including a Summary page, a new Gene Ontology page, and a new Literature page for each complex. Just as we do for all of your favorite genes, Gene Ontology and Literature curation for complexes will be ongoing.

If you have any questions or feedback about the updates to our complex pages, please do not hesitate to contact us at any time.
Categories: Announcements, Data updates, Website changes